Wittmann Lab
Research Publications Group members BiographyProtein disorder in transcription
Transcription is a fundamental process important for all living things. It needs to be precisely executed for cells to function properly during normal growth and in times of stress. If this regulation is compromised the transcriptional profile is altered, which is a hallmark of cancer and occurs as our cells age. But what fascinates us most is how transcriptional programmes can be controlled so accurately in time and space that entire organisms can develop. Here, transcription factors with so-called pioneering activity determine the cell type by promoting the expression of specific genes that govern cellular identity. During development, initially totipotent cells progressively specialise more and more in a stepwise manner. In the laboratory, we are also able to go in the opposite direction and reprogram specialised somatic cells back to a pluripotent state.
Gene activation
In either case, activation of many genes is needed to push a cell in a new direction. Before a gene can become active, however, it first needs to be found in the vast genome of the human cell. This is primarily the responsibility of transcription factors that scan the DNA for regulatory regions containing clusters of binding sites. After recognition of the region, hundreds of downstream factors are recruited and cooperate to open the chromatin, initiate transcription and determine the activity level of the gene(s).
Many decades of intricate biochemical and structural work have elucidated how the transcriptional machinery synthesises RNA and which proteins participate in the process. However, the regulation of transcription is still only poorly understood. Interestingly, proteins involved in transcription have the highest percentage of disordered content in the entire proteome because they contain the most intrinsically disordered regions (IDRs). These IDRs do not fold into a pre-defined three-dimensional structure, like enzymatic regions, but instead remain flexible and dynamically change their conformation. Using these regions, transcriptional proteins are able to interact and communicate with one another.
In our group, we are trying to understand how IDRs are used for communication and transcriptional regulation. One of their functions probably lies in the formation of liquid droplets (see below). In addition, they contain binding sites for other factors that are used for recruitment. In our group, we are trying to dissect which properties of the IDR determine which function. Are the same regions used for protein:protein interactions and droplet formation, or is there a separation of function? How does droplet formation influence DNA binding and sequence recognition? And do droplets have an advantage for transcription?
Phase transition
Recently, we and others were able to show that when transcription factors come together on DNA they can form a liquid droplet that de-mixes from the surrounding medium, thereby transitioning from a diluted state into a separate liquid phase (Figure 1). Our previous work suggested that a pre-wetting-like transition is at play here (Morin et al, Nature Physics, 2022): many transcription factors bind to clusters of binding sites on the DNA that are in close proximity. This locally increases the protein concentration. As a result, the IDRs can now interact with each other strongly enough to form a condensate or liquid droplet on the DNA. Interestingly, the resulting droplets are limited in size and restricted to the DNA surface.
While these results are intriguing, this phenomenon has so far only been studied in vitro. This is why the exact nature of transcriptional droplets inside cells remains a matter of scientific debate. One of our research projects is trying to investigate this exact topic. How many molecules come together? What is their arrangement inside the droplet? And which physical description most accurately describes what we observe in vivo?
How we work
Our group is trying to better understand how transcriptional droplets form, which proteins they contain and how they are used to control transcriptional programs. To this end, we are using a multi-disciplinary approach that involves biochemistry, cellular biology and close collaborations with biophysicists.
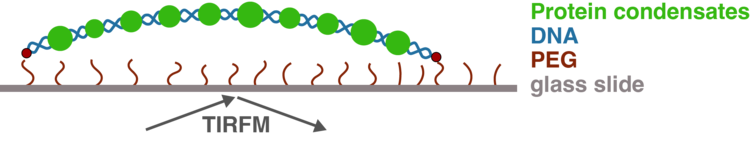
Specifically, we use recombinant proteins to test their ability to form condensates in vitro in the presence and absence of DNA. Using single-molecule assays (Figure 2), we can probe the properties of the condensates and biophysically characterise them. By creating different mutants, we can test which parts of the protein are required for different functions or are responsible for which property. Finally, we can introduce mutations into human cells to understand how they affect transcription using state-of-the-art next-generation sequencing methods. One of the proteins we use as a model is the transcription factor Klf4, which is not only important because of its role in differentiation and reprogramming, but was shown to help rejuvenate aged cells. By combining all these different techniques, we can look at transcription factors from many different angles and paint a comprehensive picture of how they function inside cells.
Recruiting
We are always happy to welcome motivated new people to our international team.
Postdoc applicants are encouraged to contact us by email.
PhD applicants can apply through the International PhD Programme. Any projects offered are listed on the website.
Undergraduate students can inquire about internship opportunities or thesis projects via email or apply directly through the IMB Internship Programme.